Efficient Tabular Storage
tl;dr: We discuss efficient techniques for on-disk storage of tabular data, notably the following:
- Binary stores
- Column stores
- Categorical support
- Compression
- Indexed/Partitioned stores
We use NYCTaxi dataset for examples, and introduce a small project, Castra.
This work is supported by Continuum Analytics and the XDATA Program as part of the Blaze Project
Larger than Memory Data and Disk I/O
We analyze large datasets (10-100GB) on our laptop by extending memory with disk. Tools like dask.array and dask.dataframe make this easier for array and tabular data.
Interaction times can improve significantly (from minutes to seconds) if we choose to store our data on disk efficiently. This is particularly important for large data because we can no longer separately “load in our data” while we get a coffee and then iterate rapidly on our dataset once it’s comfortably in memory.
Larger-than-memory datasets force interactive workflows to include the hard drive.
CSV is convenient but slow
CSV is great. It’s human readable, accessible by every tool (even Excel!), and pretty simple.
CSV is also slow. The pandas.read_csv parser maxes out at 100MB/s
on simple data. This doesn’t include any keyword arguments like datetime
parsing that might slow it down further. Consider the time to parse a 24GB
dataset:
24GB / (100MB/s) == 4 minutes
A four minute delay is too long for interactivity. We need to operate in seconds rather than minutes otherwise people leave to work on something else. This improvement from a few minutes to a few seconds is entirely possible if we choose better formats.
Example with CSVs
As an example lets play with the NYC Taxi dataset using dask.dataframe, a library that copies the Pandas API but operates in chunks off of disk.
>>> import dask.dataframe as dd
>>> df = dd.read_csv('csv/trip_data_*.csv',
... skipinitialspace=True,
... parse_dates=['pickup_datetime', 'dropoff_datetime'])
>>> df.head()| medallion | hack_license | vendor_id | rate_code | store_and_fwd_flag | pickup_datetime | dropoff_datetime | passenger_count | trip_time_in_secs | trip_distance | pickup_longitude | pickup_latitude | dropoff_longitude | dropoff_latitude | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 89D227B655E5C82AECF13C3F540D4CF4 | BA96DE419E711691B9445D6A6307C170 | CMT | 1 | N | 2013-01-01 15:11:48 | 2013-01-01 15:18:10 | 4 | 382 | 1.0 | -73.978165 | 40.757977 | -73.989838 | 40.751171 |
| 1 | 0BD7C8F5BA12B88E0B67BED28BEA73D8 | 9FD8F69F0804BDB5549F40E9DA1BE472 | CMT | 1 | N | 2013-01-06 00:18:35 | 2013-01-06 00:22:54 | 1 | 259 | 1.5 | -74.006683 | 40.731781 | -73.994499 | 40.750660 |
| 2 | 0BD7C8F5BA12B88E0B67BED28BEA73D8 | 9FD8F69F0804BDB5549F40E9DA1BE472 | CMT | 1 | N | 2013-01-05 18:49:41 | 2013-01-05 18:54:23 | 1 | 282 | 1.1 | -74.004707 | 40.737770 | -74.009834 | 40.726002 |
| 3 | DFD2202EE08F7A8DC9A57B02ACB81FE2 | 51EE87E3205C985EF8431D850C786310 | CMT | 1 | N | 2013-01-07 23:54:15 | 2013-01-07 23:58:20 | 2 | 244 | 0.7 | -73.974602 | 40.759945 | -73.984734 | 40.759388 |
| 4 | DFD2202EE08F7A8DC9A57B02ACB81FE2 | 51EE87E3205C985EF8431D850C786310 | CMT | 1 | N | 2013-01-07 23:25:03 | 2013-01-07 23:34:24 | 1 | 560 | 2.1 | -73.976250 | 40.748528 | -74.002586 | 40.747868 |
Time Costs
It takes a second to load the first few lines but 11 to 12 minutes to roll through the entire dataset. We make a zoomable picture below of a random sample of the taxi pickup locations in New York City. This example is taken from a full example notebook here.
df2 = df[(df.pickup_latitude > 40) &
(df.pickup_latitude < 42) &
(df.pickup_longitude > -75) &
(df.pickup_longitude < -72)]
sample = df2.sample(frac=0.0001)
pickup = sample[['pickup_latitude', 'pickup_longitude']]
result = pickup.compute()
from bokeh.plotting import figure, show, output_notebook
p = figure(title="Pickup Locations")
p.scatter(result.pickup_longitude, result.pickup_latitude, size=3, alpha=0.2)Eleven minutes is a long time
This result takes eleven minutes to compute, almost all of which is parsing CSV files. While this may be acceptable for a single computation we invariably make mistakes and start over or find new avenues in our data to explore. Each step in our thought process now takes eleven minutes, ouch.
Interactive exploration of larger-than-memory datasets requires us to evolve beyond CSV files.
Principles to store tabular data
What efficient techniques exist for tabular data?
A good solution may have the following attributes:
- Binary
- Columnar
- Categorical support
- Compressed
- Indexed/Partitioned
We discuss each of these below.
Binary
Consider the text ‘1.23’ as it is stored in a CSV file and how it is stored as a Python/C float in memory:
- CSV:
1.23 - C/Python float:
0x3f9d70a4
These look very different. When we load 1.23 from a CSV textfile
we need to translate it to 0x3f9d70a4; this takes time.
A binary format stores our data on disk exactly how it will look in memory; we
store the bytes 0x3f9d70a4 directly on disk so that when we load data
from disk to memory no extra translation is necessary. Our file is no longer
human readable but it’s much faster.
This gets more intense when we consider datetimes:
- CSV: 2015-08-25 12:13:14
- NumPy datetime representation: 1440529994000000 (as an integer)
Every time we parse a datetime we need to compute how many microseconds it has
been since the epoch. This calculation needs to take into account things like
how many days in each month, and all of the intervening leap years. This is
slow. A binary representation would record the integer directly on disk (as
0x51e278694a680) so that we can load our datetimes directly into memory
without calculation.
Columnar
Many analytic computations only require a few columns at a time, often only one, e.g.
>>> df.passenger_counts.value_counts().compute().sort_index()
0 3755
1 119605039
2 23097153
3 7187354
4 3519779
5 9852539
6 6628287
7 30
8 23
9 24
129 1
255 1
Name: passenger_count, dtype: int64Of our 24 GB we may only need 2GB. Columnar storage means storing each column separately from the others so that we can read relevant columns without passing through irrelevant columns.
Our CSV example fails at this. While we only want two columns,
pickup_datetime and pickup_longitude, we pass through all of our data to
collect the relevant fields. The pickup location data is mixed with all the
rest.
Categoricals
Categoricals encode repetitive text columns (normally very expensive) as integers (very very cheap) in a way that is invisible to the user.
Consider the following (mostly text) columns of our NYC taxi dataset:
>>> df[['medallion', 'vendor_id', 'rate_code', 'store_and_fwd_flag']].head()| medallion | vendor_id | rate_code | store_and_fwd_flag | |
|---|---|---|---|---|
| 0 | 89D227B655E5C82AECF13C3F540D4CF4 | CMT | 1 | N |
| 1 | 0BD7C8F5BA12B88E0B67BED28BEA73D8 | CMT | 1 | N |
| 2 | 0BD7C8F5BA12B88E0B67BED28BEA73D8 | CMT | 1 | N |
| 3 | DFD2202EE08F7A8DC9A57B02ACB81FE2 | CMT | 1 | N |
| 4 | DFD2202EE08F7A8DC9A57B02ACB81FE2 | CMT | 1 | N |
Each of these columns represents elements of a small set:
- There are two vendor ids
- There are twenty one rate codes
- There are three store-and-forward flags (Y, N, missing)
- There are about 13000 taxi medallions. (still a small number)
And yet we store these elements in large and cumbersome dtypes:
In [4]: df[['medallion', 'vendor_id', 'rate_code', 'store_and_fwd_flag']].dtypes
Out[4]:
medallion object
vendor_id object
rate_code int64
store_and_fwd_flag object
dtype: objectWe use int64 for rate code, which could easily have fit into an int8 an
opportunity for an 8x improvement in memory use. The object dtype used for
strings in Pandas and Python takes up a lot of memory and is quite slow:
In [1]: import sys
In [2]: sys.getsizeof('CMT') # bytes
Out[2]: 40Categoricals replace the original column with a column of integers (of the
appropriate size, often int8) along with a small index mapping those integers to the
original values. I’ve written about categoricals
before so I
won’t go into too much depth here. Categoricals increase both storage and
computational efficiency by about 10x if you have text data that describes
elements in a category.
Compression
After we’ve encoded everything well and separated our columns we find ourselves limited by disk I/O read speeds. Disk read bandwidths range from 100MB/s (laptop spinning disk hard drive) to 2GB/s (RAID of SSDs). This read speed strongly depends on how large our reads are. The bandwidths given above reflect large sequential reads such as you might find when reading all of a 100MB file in one go. Performance degrades for smaller reads. Fortunately, for analytic queries we’re often in the large sequential read case (hooray!)
We reduce disk read times through compression. Consider the datetimes of the NYC taxi dataset. These values are repetitive and slowly changing; a perfect match for modern compression techniques.
>>> ind = df.index.compute() # this is on presorted index data (see castra section below)
>>> ind
DatetimeIndex(['2013-01-01 00:00:00', '2013-01-01 00:00:00',
'2013-01-01 00:00:00', '2013-01-01 00:00:00',
'2013-01-01 00:00:00', '2013-01-01 00:00:00',
'2013-01-01 00:00:00', '2013-01-01 00:00:00',
'2013-01-01 00:00:00', '2013-01-01 00:00:00',
...
'2013-12-31 23:59:42', '2013-12-31 23:59:47',
'2013-12-31 23:59:48', '2013-12-31 23:59:49',
'2013-12-31 23:59:50', '2013-12-31 23:59:51',
'2013-12-31 23:59:54', '2013-12-31 23:59:55',
'2013-12-31 23:59:57', '2013-12-31 23:59:57'],
dtype='datetime64[ns]', name=u'pickup_datetime', length=169893985, freq=None, tz=None)Benchmark datetime compression
We can use a modern compression library, like fastlz or blosc to compress this data at high speeds.
In [36]: import blosc
In [37]: %time compressed = blosc.compress_ptr(address=ind.values.ctypes.data,
...: items=len(ind),
...: typesize=ind.values.dtype.alignment,
...: clevel=5)
CPU times: user 3.22 s, sys: 332 ms, total: 3.55 s
Wall time: 512 ms
In [40]: len(compressed) / ind.nbytes # compression ratio
Out[40]: 0.14296813539337488
In [41]: ind.nbytes / 0.512 / 1e9 # Compresson bandwidth (GB/s)
Out[41]: 2.654593515625
In [42]: %time _ = blosc.decompress(compressed)
CPU times: user 1.3 s, sys: 438 ms, total: 1.74 s
Wall time: 406 ms
In [43]: ind.nbytes / 0.406 / 1e9 # Decompression bandwidth (GB/s)
Out[43]: 3.3476647290640393We store 7x fewer bytes on disk (thus septupling our effective disk I/O) by adding an extra 3GB/s delay. If we’re on a really nice Macbook pro hard drive (~600MB/s) then this is a clear and substantial win. The worse the hard drive, the better this is.
\[\textrm{Effective bandwidth} = \left(\frac{1}{600 \textrm{MB/s} \times 7} + \frac{1}{3000 \textrm{MB/s}}\right)^{-1} = 1750 \textrm{MB/s}\]But sometimes compression isn’t as nice
Some data is more or less compressable than others. The following column of floating point data does not compress as nicely.
In [44]: x = df.pickup_latitude.compute().values
In [45]: %time compressed = blosc.compress_ptr(x.ctypes.data, len(x), x.dtype.alignment, clevel=5)
CPU times: user 5.87 s, sys: 0 ns, total: 5.87 s
Wall time: 925 ms
In [46]: len(compressed) / x.nbytes
Out[46]: 0.7518617315969132This compresses more slowly and only provides marginal benefit. Compression may still be worth it on slow disk but this isn’t a huge win.
The pickup_latitude column isn’t compressible because most of the information isn’t repetitive. The numbers to the far right of the decimal point are more or less random.
40.747868
Other floating point columns may compress well, particularly when they are rounded to small and meaningful decimal values.
Compression rules of thumb
Optimal compression requires thought. General rules of thumb include the following:
- Compress integer dtypes
- Compress datetimes
- If your data is slowly varying (e.g. sorted time series) then use a shuffle filter (default in blosc)
- Don’t bother much with floating point dtypes
- Compress categoricals (which are just integer dtypes)
Avoid gzip and bz2
Finally, avoid gzip and bz2. These are both very common and very slow. If dealing with text data, consider snappy (also available via blosc.)
Indexing/Partitioning
One column usually dominates our queries. In time-series data this is time. For personal data this is the user ID.
Just as column stores let us avoid irrelevant columns, partitioning our data along a preferred index column lets us avoid irrelevant rows. We may need the data for the last month and don’t need several years’ worth. We may need the information for Alice and don’t need the information for Bob.
Traditional relational databases provide indexes on any number of columns or sets of columns. This is excellent if you are using a traditional relational database. Unfortunately the data structures to provide arbitrary indexes don’t mix well with some of the attributes discussed above and we’re limited to a single index that partitions our data into sorted blocks.
Some projects that implement these principles
Many modern distributed database storage systems designed for analytic queries implement these principles well. Notable players include Redshift and Parquet.
Additionally newer single-machine data stores like Dato’s SFrame and BColz follow many of these principles. Finally many people have been doing this for a long time with custom use of libraries like HDF5.
It turns out that these principles are actually quite easy to implement with the right tools (thank you #PyData) The rest of this post will talk about a tiny 500 line project, Castra, that implements these princples and gets good speedups on biggish Pandas data.
Castra
With these goals in mind we built Castra, a binary partitioned compressed columnstore with builtin support for categoricals and integration with both Pandas and dask.dataframe.
Load data from CSV files, sort on index, save to Castra
Here we load in our data from CSV files, sort on the pickup datetime column, and store to a castra file. This takes about an hour (as compared to eleven minutes for a single read.) Again, you can view the full notebook here
>>> import dask.dataframe as dd
>>> df = dd.read_csv('csv/trip_data_*.csv',
... skipinitialspace=True,
... parse_dates=['pickup_datetime', 'dropoff_datetime'])
>>> (df.set_index('pickup_datetime', compute=False)
... .to_castra('trip.castra', categories=True))Profit
Now we can take advantage of columnstores, compression, and binary representation to perform analytic queries quickly. Here is code to create a histogram of trip distance. The plot of the results follows below.
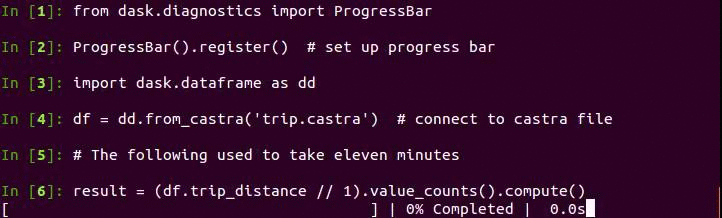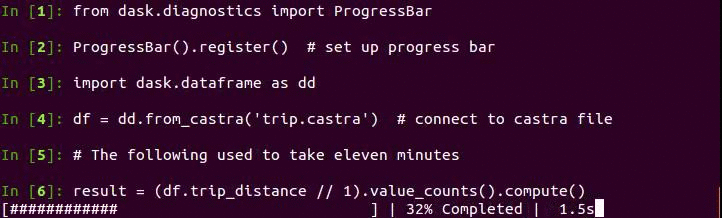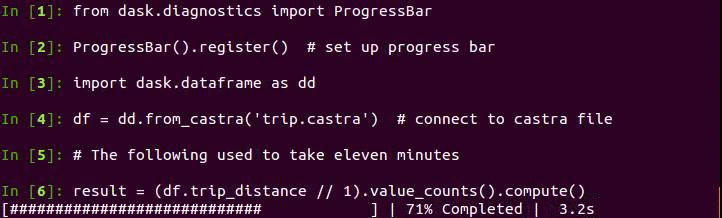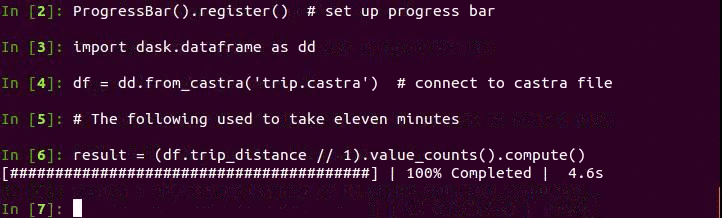
Note that this is especially fast because Pandas now releases the
GIL on value_counts
operations (all groupby operations really). This takes around 20 seconds on
my machine on the last release of Pandas vs 5 seconds on the development
branch. Moving from CSV files to Castra moved the bottleneck of our
computation from disk I/O to processing speed, allowing improvements like
multi-core processing to really shine.
We plot the result of the above computation with Bokeh below. Note the spike around 20km. This is around the distance from Midtown Manhattan to LaGuardia airport.
I’ve shown Castra used above with dask.dataframe but it works fine with straight Pandas too.
Credit
Castra was started by myself and Valentin Haenel (current maintainer of bloscpack and bcolz) during an evening sprint following PyData Berlin. Several bugfixes and refactors were followed up by Phil Cloud and Jim Crist.
Castra is roughly 500 lines long. It’s a tiny project which is both good and bad. It’s being used experimentally and there are some heavy disclaimers in the README. This post is not intended as a sales pitch for Castra, but rather to provide a vocabulary to talk about efficient tabular storage.
Response to twitter traffic: again, this blogpost is not saying “use Castra!” Rather it says “don’t use CSVs!” and consider more efficient storage for interactive use. Other, more mature solutions exist or could be built. Castra was an experiment of “how fast can we get without doing too much work.”
blog comments powered by Disqus