Custom Parallel Algorithms on a Cluster with Dask
This work is supported by Continuum Analytics the XDATA Program and the Data Driven Discovery Initiative from the Moore Foundation
Summary
This post describes Dask as a computational task scheduler that fits somewhere on a spectrum between big data computing frameworks like Hadoop/Spark and task schedulers like Airflow/Celery/Luigi. We see how, by combining elements from both of these types of systems Dask is able to handle complex data science problems particularly well.
This post is in contrast to two recent posts on structured parallel collections:
Big Data Collections
Most distributed computing systems like Hadoop or Spark or SQL databases implement a small but powerful set of parallel operations like map, reduce, groupby, and join. As long as you write your programs using only those operations then the platforms understand your program and serve you well. Most of the time this is great because most big data problems are pretty simple.
However, as we explore new complex algorithms or messier data science problems, these large parallel operations start to become insufficiently flexible. For example, consider the following data loading and cleaning problem:
- Load data from 100 different files (this is a simple
mapoperation) - Also load a reference dataset from a SQL database (not parallel at all, but could run alongside the map above)
- Normalize each of the 100 datasets against the reference dataset (sort of like a map, but with another input)
- Consider a sliding window of every three normalized datasets (Might be able to hack this with a very clever join? Not sure.)
- Of all of the 98 outputs of the last stage, consider all pairs. (Join or cartesian product) However, because we don’t want to compute all ~10000 possibilities, let’s just evaluate a random sample of these pairs
- Find the best of all of these possibilities (reduction)
In sequential for-loopy code this might look like the following:
filenames = ['mydata-%d.dat' % i for i in range(100)]
data = [load(fn) for fn in filenames]
reference = load_from_sql('sql://mytable')
processed = [process(d, reference) for d in data]
rolled = []
for i in range(len(processed) - 2):
a = processed[i]
b = processed[i + 1]
c = processed[i + 2]
r = roll(a, b, c)
rolled.append(r)
compared = []
for i in range(200):
a = random.choice(rolled)
b = random.choice(rolled)
c = compare(a, b)
compared.append(c)
best = reduction(compared)
This code is clearly parallelizeable, but it’s not clear how to write it down as a MapReduce program, a Spark computation, or a SQL query. These tools generally fail when asked to express complex or messy problems. We can still use Hadoop/Spark to solve this problem, but we are often forced to change and simplify our objectives a bit. (This problem is not particularly complex, and I suspect that there are clever ways to do it, but it’s not trivial and often inefficient.)
Task Schedulers
So instead people use task schedulers like Celery, Luigi, or Airflow. These systems track hundreds of tasks, each of which is just a normal Python function that runs on some normal Python data. The task scheduler tracks dependencies between tasks and so runs as many as it can at once if they don’t depend on each other.
This is a far more granular approach than the Big-Bulk-Collection approach of MapReduce and Spark. However systems like Celery, Luigi, and Airflow are also generally less efficient. This is both because they know less about their computations (map is much easier to schedule than an arbitrary graph) and because they just don’t have machinery for inter-worker communication, efficient serialization of custom datatypes, etc..
Dask Mixes Task Scheduling with Efficient Computation
Dask is both a big data system like Hadoop/Spark that is aware of resilience, inter-worker communication, live state, etc. and also a general task scheduler like Celery, Luigi, or Airflow, capable of arbitrary task execution.
Many Dask users use something like Dask dataframe, which generates these graphs automatically, and so never really observe the task scheduler aspect of Dask This is, however, the core of what distinguishes Dask from other systems like Hadoop and Spark. Dask is incredibly flexible in the kinds of algorithms it can run. This is because, at its core, it can run any graph of tasks and not just map, reduce, groupby, join, etc.. Users can do this natively, without having to subclass anything or extend Dask to get this extra power.
There are significant performance advantages to this. For example:
- Dask.dataframe can easily represent nearest neighbor computations for fast time-series algorithms
- Dask.array can implement complex linear algebra solvers or SVD algorithms from the latest research
- Complex Machine Learning algorithms are often easier to implement in Dask, allowing it to be more efficient through smarter algorithms, as well as through scalable computing.
- Complex hierarchies from bespoke data storage solutions can be explicitly modeled and loaded in to other Dask systems
This doesn’t come for free. Dask’s scheduler has to be very intelligent to smoothly schedule arbitrary graphs while still optimizing for data locality, worker failure, minimal communication, load balancing, scarce resources like GPUs and more. It’s a tough job.
Dask.delayed
So let’s go ahead and run the data ingestion job described with Dask.
We craft some fake functions to simulate actual work:
import random
from time import sleep
def load(address):
sleep(random.random() / 2)
def load_from_sql(address):
sleep(random.random() / 2 + 0.5)
def process(data, reference):
sleep(random.random() / 2)
def roll(a, b, c):
sleep(random.random() / 5)
def compare(a, b):
sleep(random.random() / 10)
def reduction(seq):
sleep(random.random() / 1)
We annotate these functions with dask.delayed, which changes a function so
that instead of running immediately it captures its inputs and puts everything
into a task graph for future execution.
from dask import delayed
load = delayed(load)
load_from_sql = delayed(load_from_sql)
process = delayed(process)
roll = delayed(roll)
compare = delayed(compare)
reduction = delayed(reduction)
Now we just call our normal Python for-loopy code from before. However now rather than run immediately our functions capture a computational graph that can be run elsewhere.
filenames = ['mydata-%d.dat' % i for i in range(100)]
data = [load(fn) for fn in filenames]
reference = load_from_sql('sql://mytable')
processed = [process(d, reference) for d in data]
rolled = []
for i in range(len(processed) - 2):
a = processed[i]
b = processed[i + 1]
c = processed[i + 2]
r = roll(a, b, c)
rolled.append(r)
compared = []
for i in range(200):
a = random.choice(rolled)
b = random.choice(rolled)
c = compare(a, b)
compared.append(c)
best = reduction(compared)
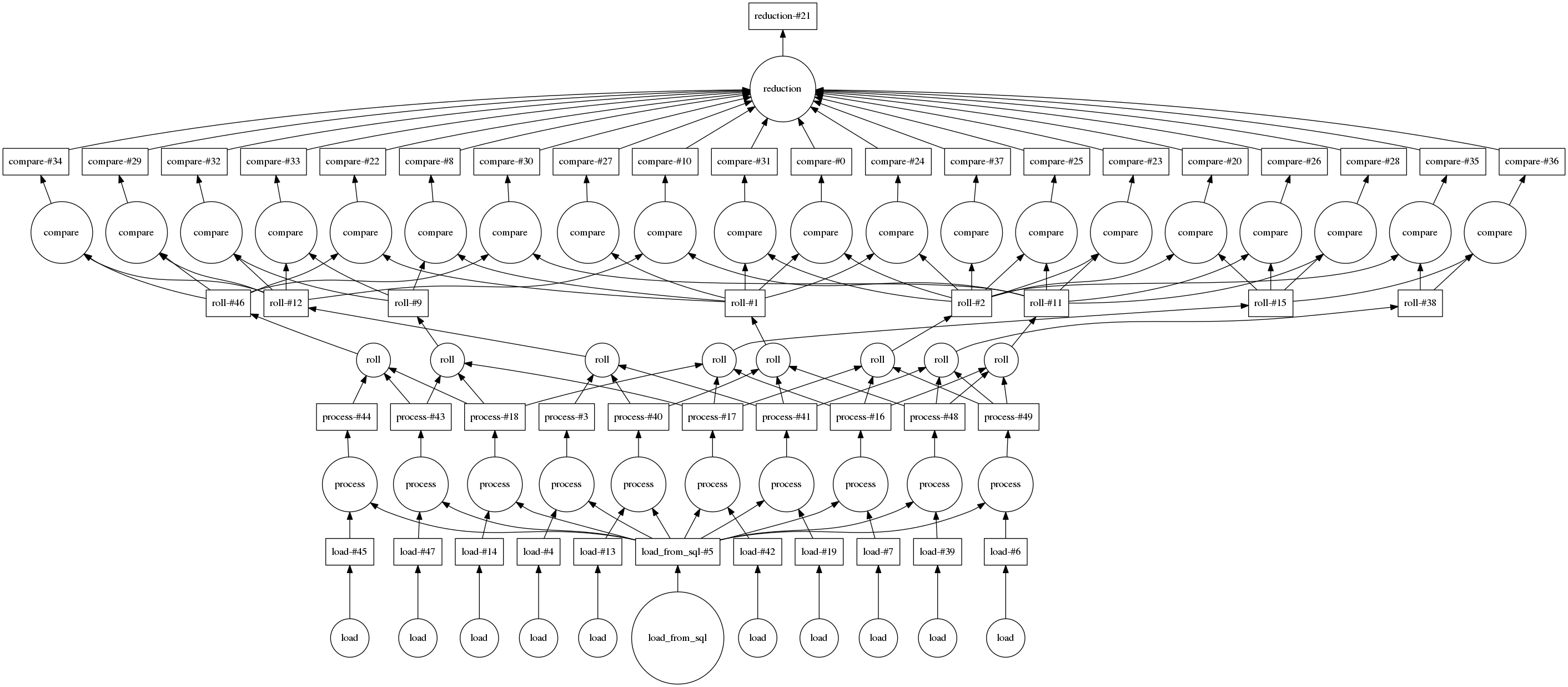
Here is an image of that graph for a smaller input of only 10 files and 20 random pairs
We can connect to a small cluster with 20 cores
from dask.distributed import Client
client = Client('scheduler-address:8786')
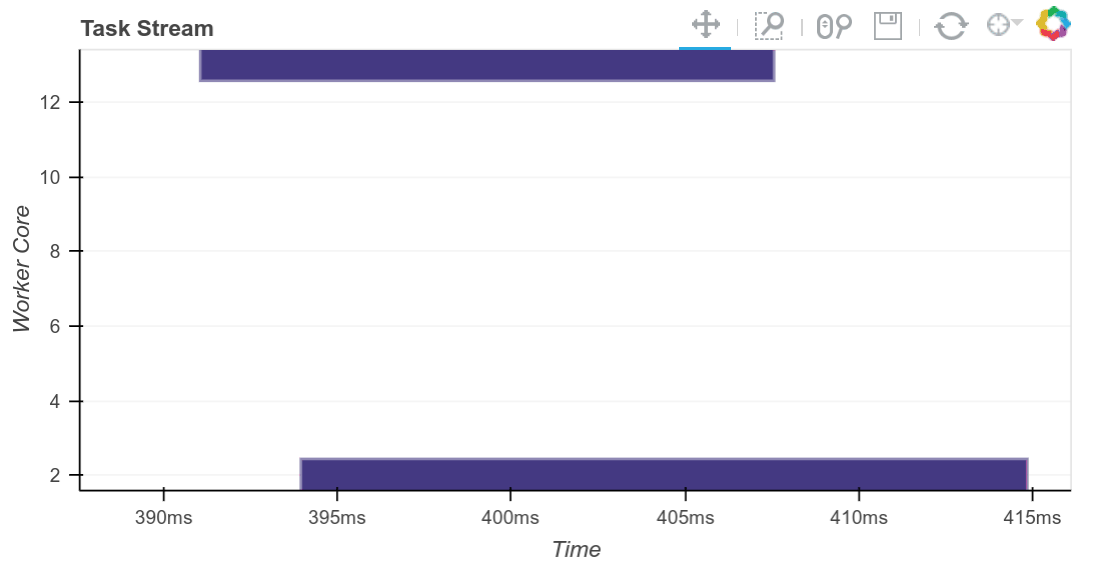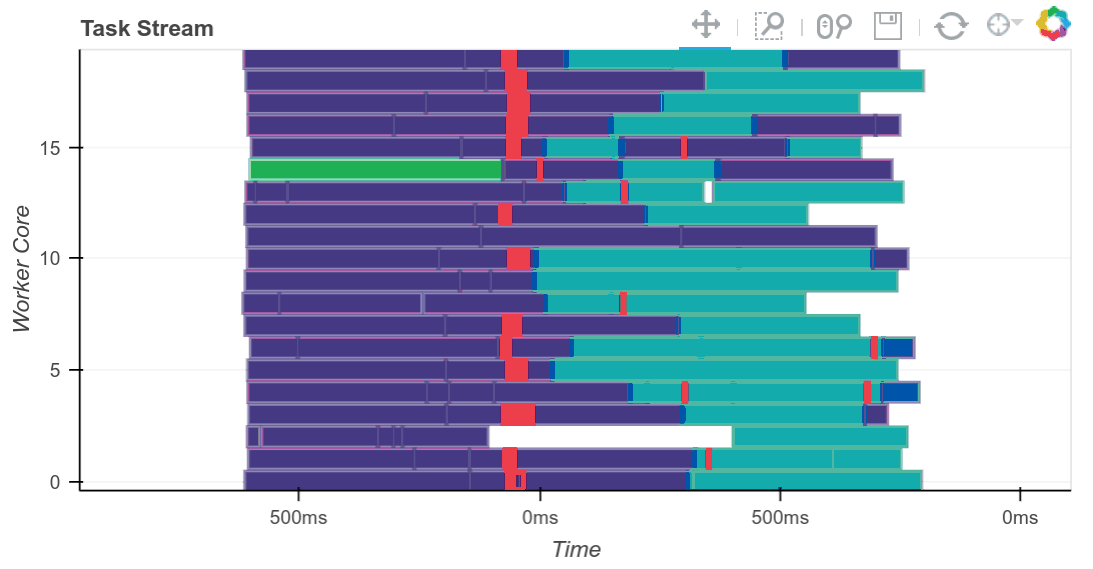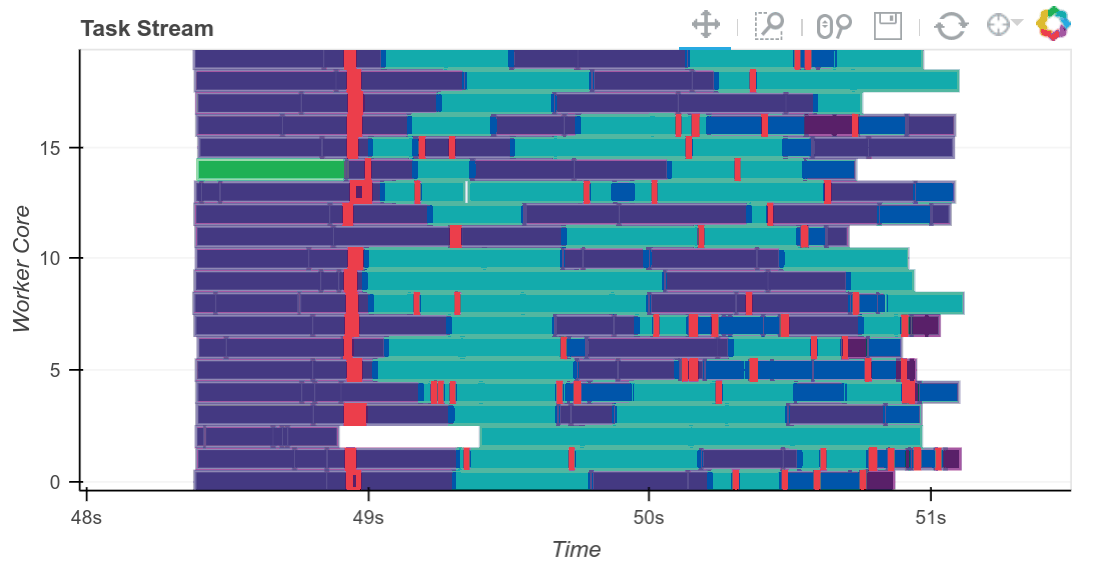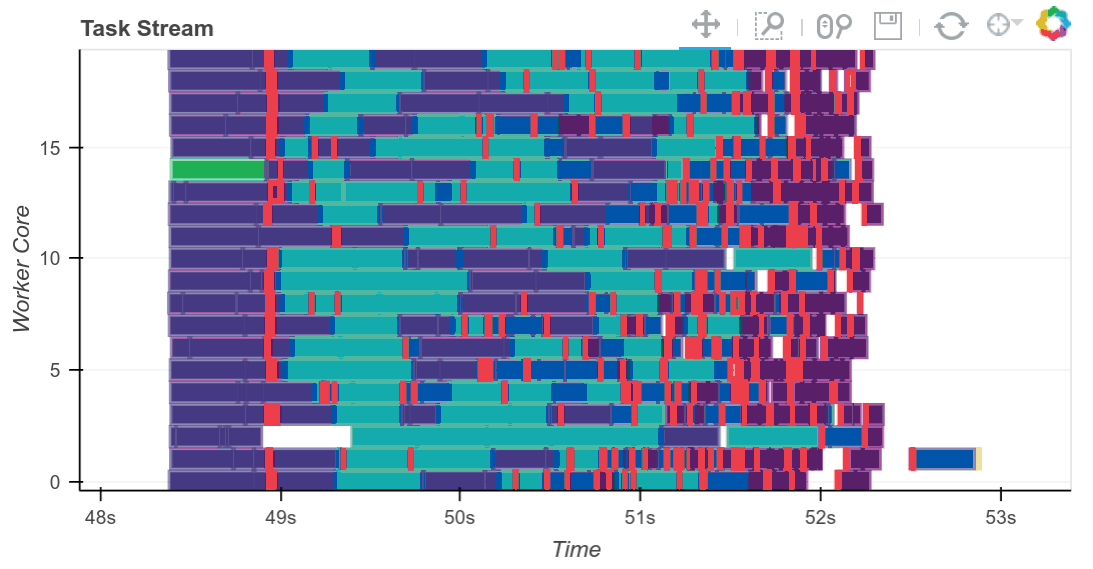
We compute the result and see the trace of the computation running in real time.
result = best.compute()
The completed Bokeh image below is interactive. You can pan and zoom by selecting the tools in the upper right. You can see every task, which worker it ran on and how long it took by hovering over the rectangles.
We see that we use all 20 cores well. Intermediate results are transferred between workers as necessary (these are the red rectangles). We can scale this up as necessary. Dask scales to thousands of cores.
Final Thoughts
Dask’s ability to write down arbitrary computational graphs Celery/Luigi/Airflow-style and yet run them with the scalability promises of Hadoop/Spark allows for a pleasant freedom to write comfortably and yet still compute scalably. This ability opens up new possibilities both to support more sophisticated algorithms and also to handle messy situations that arise in the real world (enterprise data systems are sometimes messy) while still remaining within the bounds of “normal and supported” Dask operation.
blog comments powered by Disqus